OpenFold3 Meets AMD Instinct™ GPUs: Unlocking Scalable, High-Throughput Structural Biology
Apr 09, 2026
Biology is entering an inference-at-scale era.
A decade ago, accurately predicting the structure of a biological molecule from sequence alone seemed out of reach. Today, it is a reality. Following the breakthrough of AlphaFold2, a new generation of machine learning models like AlphaFold3 can predict not just single protein structures, but entire biological complexes: proteins bound to ligands, antibodies interacting with antigens, and transcription factors engaging DNA. This shift matters because biology does not happen in isolation. Proteins, nucleic acids, ligands, ions, sugars, and lipids interact in dense, dynamic environments. To study disease, develop therapeutics, and engineer new biology, researchers need tools that can capture these interactions quickly, accurately, and at scale.
That is where OpenFold3 and AMD Instinct™ GPUs come together.
OpenFold3 builds on the foundation of AlphaFold3 and brings advanced co-folding capabilities into a truly open ecosystem, giving researchers and industry access to open code, open weights, and open workflows for studying complex biological systems. AMD Instinct™ GPUs provide the performance, memory capacity, and scalability needed to run these models efficiently on real workloads. Together, OpenFold3 and AMD Instinct™ GPUs move structural biology from isolated prediction to scalable discovery.
Fast Inference and Large-Context Scaling on AMD Instinct™ GPUs
Co-folding is a major leap in biological modeling, but it comes with steep computational costs. As sequence length increases and molecular context becomes richer, OpenFold3 inference becomes more computationally intensive. For real-world workflows such as protein–ligand screening, antibody candidate selection, and large-complex modeling, both speed and memory become critical.
In close collaboration with the OpenFold3 team and Prof. Minjia Zhang’s group at the University of Illinois Urbana-Champaign, AMD accelerated OpenFold3 inference using hardware-agnostic Triton kernels to bring memory-efficient OpenFold3 inference across a broader range of hardware platforms.
The results on GPU-based test systems are strong: up to 45.5% faster inference performance on an AMD Instinct™ MI355X versus NVIDIA B200 at sequence length 5,000, and up to 22.1% faster inference on AMD Instinct™ MI300X, versus NVIDIA H100 at sequence length 3,0002.
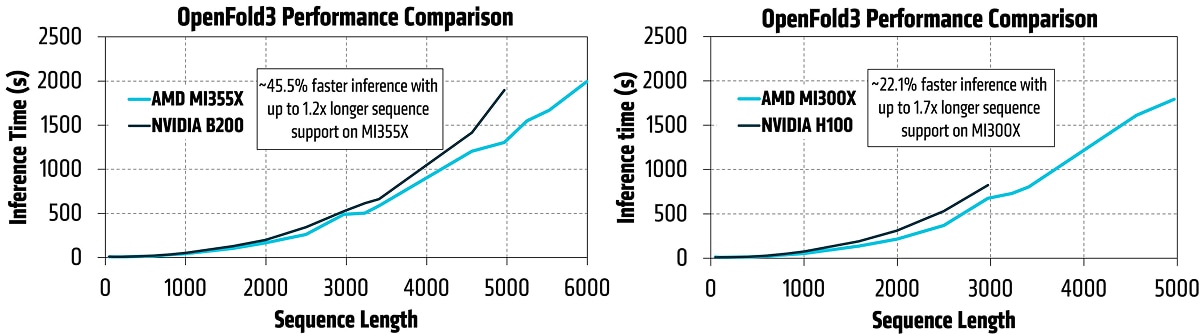
AMD Instinct™ GPUs also extend the scale of biomolecular workloads. Co-folding models rely on dense pair representations whose memory footprint grows quadratically with the number of tokens, making memory capacity a key bottleneck as input size grows. That has historically constrained many methods to moderate sequence lengths, even when the biology requires a much larger context.
The AMD memory advantage helps remove that constraint: in the AMD benchmarking, the MI355X GPU supported sequence lengths up to 6,000, while the NVIDIA B200 ran out of memory at 5,000. Similarly, the MI300X GPU reached 5,000, while the NVIDIA H100 ran out of memory at 3,000. This added memory headroom enables larger and richer biomolecular modeling with fewer compromises in fidelity.
Make It Open. Make It Fast. Make It Scalable.
Built for the next scale of structural biology.
As biomolecular foundation models become more capable, the bottleneck is no longer just model quality. What matters now is whether these systems can run fast enough, scale far enough, and remain open enough to drive real-world discovery.
AMD Instinct™ GPUs provide the memory bandwidth and capacity required to run long-context workloads at scale.
This is the inflection point: moving long-context structural prediction from isolated demonstrations to sustained, production-grade throughput. It enables researchers and industry to tackle real biological complexity with fewer compromises and faster iteration toward new therapeutics and precision medicine.
Explore OpenFold3 on AMD Instinct™ GPUs
Interested in running OpenFold3 on AMD Instinct™ GPUs?
Check out the latest OpenFold3 release for easier installation and deployment. Visit the OpenFold3 project website for code, documentation, and usage guidance.
As the ecosystem advances, OpenFold3 on AMD Instinct™ GPUs will continue to expand what researchers can do in biomolecular modeling, therapeutic discovery, and precision medicine.
Footnotes
- Based on AMD internal testing as of March 31, 2026, using OpenFold3. Inference performance was measured at a sequence length of 5,000, the longest sequence length supported by both platforms. Maximum supported sequence length was determined by incrementally increasing sequence length until GPU memory was exhausted, reaching 6,000 on AMD and 5,000 on NVIDIA. AMD test system: AMD Instinct MI355X GPU (288 GB), AMD EPYC 9575F 64-core CPU, 3 TB RAM, ROCm 7.2. NVIDIA test system: NVIDIA B200 GPU (180 GB), Intel Xeon Platinum 8592+ CPU, 354 GB RAM, CUDA 12.9. System configurations may vary; results may differ. (MI350-076).
- Based on AMD internal testing as of March 31, 2026, using OpenFold3. Inference performance was measured at a sequence length of 3,000, the longest sequence length supported by both platforms. Maximum supported sequence length was determined by incrementally increasing sequence length until GPU memory was exhausted, reaching 5,000 on AMD and 3,000 on NVIDIA. AMD test system: AMD Instinct MI300X GPU (192 GB), AMD EPYC 9684X 96-core CPU, 2 TB RAM, ROCm 7.2. NVIDIA test system: NVIDIA H100 GPU (80 GB), AMD EPYC 9554 64-core CPU, 1.1 TB RAM, CUDA 12.9. System configurations may vary; results may differ. (MI350-077).
- Based on AMD internal testing as of March 31, 2026, using OpenFold3. Inference performance was measured at a sequence length of 5,000, the longest sequence length supported by both platforms. Maximum supported sequence length was determined by incrementally increasing sequence length until GPU memory was exhausted, reaching 6,000 on AMD and 5,000 on NVIDIA. AMD test system: AMD Instinct MI355X GPU (288 GB), AMD EPYC 9575F 64-core CPU, 3 TB RAM, ROCm 7.2. NVIDIA test system: NVIDIA B200 GPU (180 GB), Intel Xeon Platinum 8592+ CPU, 354 GB RAM, CUDA 12.9. System configurations may vary; results may differ. (MI350-076).
- Based on AMD internal testing as of March 31, 2026, using OpenFold3. Inference performance was measured at a sequence length of 3,000, the longest sequence length supported by both platforms. Maximum supported sequence length was determined by incrementally increasing sequence length until GPU memory was exhausted, reaching 5,000 on AMD and 3,000 on NVIDIA. AMD test system: AMD Instinct MI300X GPU (192 GB), AMD EPYC 9684X 96-core CPU, 2 TB RAM, ROCm 7.2. NVIDIA test system: NVIDIA H100 GPU (80 GB), AMD EPYC 9554 64-core CPU, 1.1 TB RAM, CUDA 12.9. System configurations may vary; results may differ. (MI350-077).









