Dayhoff Health, AMD Accelerate Genomic Analysis by up to 330x
Apr 09, 2026

Over the last 18 years, there’s been a fundamental transformation in genetic sequencing. The adoption of highly parallel next-generation sequencing systems back in 2008 ushered in an era of rapidly declining costs and rising performance. Over time, this has changed the bottlenecks many labs contend with.
Today, the genetic sequencer is no longer one of the slowest parts of the process. Now, secondary analysis, including alignment, classification, and matrix construction more often constrain performance. These workloads can be parallelized and scaled across CPUs, but they are memory and compute-intensive and do not scale particularly well.
The cost, memory, and compute requirements of CPU-based scaling has prompted geneticists to evaluate running certain tasks on GPUs, but any serious effort to move away from decades of CPU-based processing requires a mature software platform, validated accuracy compared to conventional analysis, broad library support, and a robust hardware architecture. While GPUs are well-suited to highly parallel processing, they dispatch and execute code very differently than a CPU and are subject to their own performance constraints.
A recent whitepaper by Dayhoff Health evaluates CPU vs. GPU processing times in genomic workloads and measures how much additional time CPU-only analysis requires compared to running the same task on a GPU. Dayhoff Health powers a clinical microbiome and genomics platform that processes high volumes of health-related data for healthcare systems and enterprise programs. The workloads evaluated in this whitepaper comprise active production pipelines that support bulk microbiome testing at scale. The company compared three types of tests: Whole genome sequencing (WGS), single-cell sequencing (SCS), and microbiome sample analysis.
For those of you (by which I mean "us") who aren't scientists, the paper explains what these three genetic tests are used for and what the benefits of accelerating them would be. Whole genome sequencing reads all the DNA in an organism's genome. It is used to detect certain rare diseases and inherited disorders, and can help oncologists tailor cancer treatment more precisely. Single-cell sequencing is used to identify and map differences between individual cells at a fine-grained level of detail. It's used in a variety of fields, including cancer research, immunology, developmental biology, neuroscience, and drug discovery.
Microbiome analysis is used to, in Dayhoff's words, "characterize microbial communities present in clinical, environmental, and research samples." A microbiome refers to a community of organisms that can be found living together in the same environment, but the term is sometimes colloquially used to refer to microorganisms commonly found living within the human gastrointestinal tract. Microbiome analysis can help physicians assess overall gut health, detect pathogens, and support various diagnostic and treatment-related decisions.
Each of these analyses benefits from substantially shorter processing times. Reducing bottlenecks in genetic analysis helps both individual patients and service providers. Individual patients can receive results more quickly, reducing potentially critical delays, while genetic testing firms can process more data in the same amount of time, improving throughput. Improving the speed and accuracy of genetic testing has been key to reducing its cost according to Genome.gov, and GPU acceleration represents the next logical step in this process.
For its CPU vs. GPU experiment, Dayhoff Health evaluated an AMD Radeon™ AI PRO R9700 graphics card with 32GB of VRAM and compared it to a 16-core AMD Ryzen™ 7950X CPU backed up by 64GB of main memory. The AMD Radeon GPU leveraged the AMD ROCm™ 7.0.2 software platform and a HIP-optimized genomics kernel; compilation flags are available on page four of the report.
While all three results are impressive, the microbiome sample analysis is particularly eye-opening. In this test, the AMD Radeon AI PRO R9700 is no less than 330x faster than its unoptimized CPU challenger, as shown below:
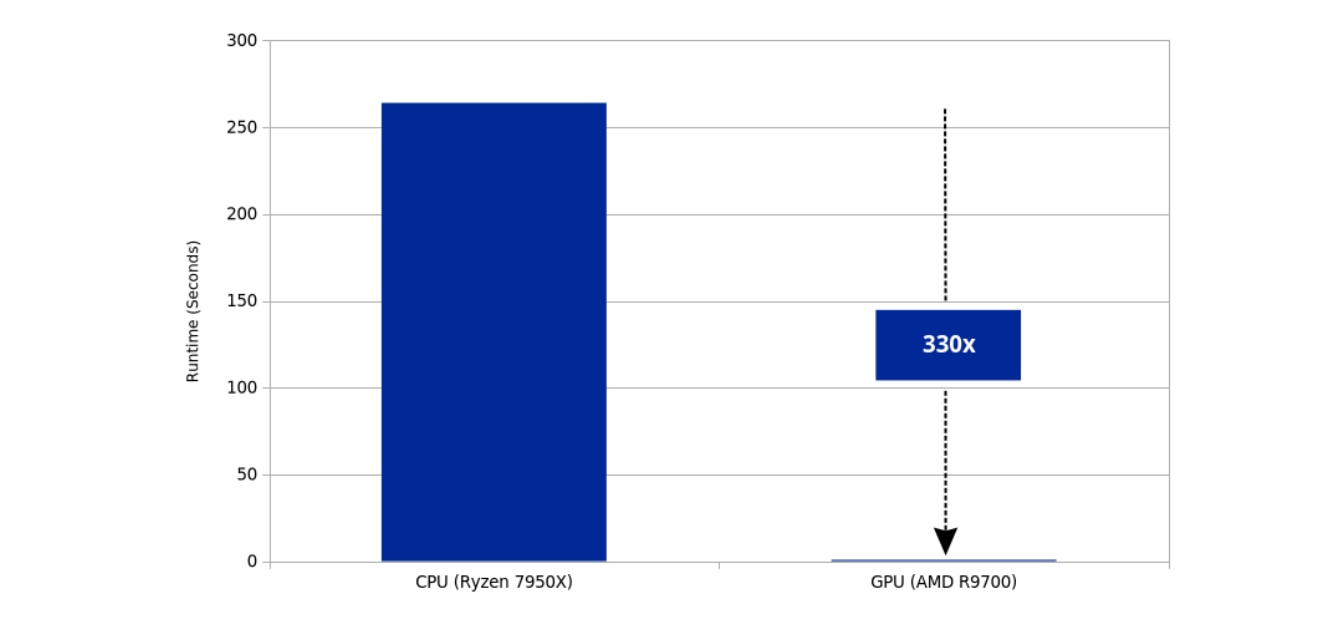
A 330× increase in microbiome pipeline processing speed reduces the time required to generate and deliver report results, bringing turnaround close to real-time. Faster processing increases overall throughput and allows large programs to serve more participants within existing infrastructure capacity. Reduced per-sample computation time improves operational efficiency and lowers per-test cost for high-volume administrators. At scale, performance gains of this magnitude enable a more responsive, economically sustainable model for microbiome and population health programs.
Optimized CPU code narrows the performance gap between CPUs and GPUs, but not enough to eliminate it. Dayhoff Health reports the GPU as ~62.5x faster than the fastest optimized CPU run. Memory pressure can degrade the situation even more; the AMD Radeon AI PRO R9700 is no less than 968x faster than the AMD Ryzen 7950X in this scenario1.
While the gains for WGS and SCS are more modest, these two workloads still report a 15x and 21x improvement for GPU processing over CPU processing under a 99% accuracy standard1. Performance gains of this magnitude are rare in modern computing, but they speak well of the AMD ROCm™ software platform, HIP-accelerated computing, and the underlying AMD RDNA™ 4 GPU microarchitecture powering the AMD Radeon AI PRO R9700.
In highly parallel genomic workloads, modern GPUs offer a structural advantage over traditional CPU architectures because they enjoy a high-throughput pool of dedicated VRAM and field thousands of cores optimized for performing identical operations across the entire chip. Dayhoff Health's findings suggest GPU acceleration will be ever-more important to DNA sequencing as its use proliferates and costs continue to drop, emphasizing the combined value and maturity of the AMD software + hardware stack.
1: GD-181a: Performance claims are provided by the 3rd party organization featured herein and have not been independently verified by AMD. Performance and cost benefits are impacted by a variety of variables. Results herein are specific to such 3rd party organization and may not be typical. GD-181a.








